Linear Regression plot with normal curves for error (sideways)
[caption id="attachment\_755" align="aligncenter" width="480" caption="Linear Regression: Normal Errors"]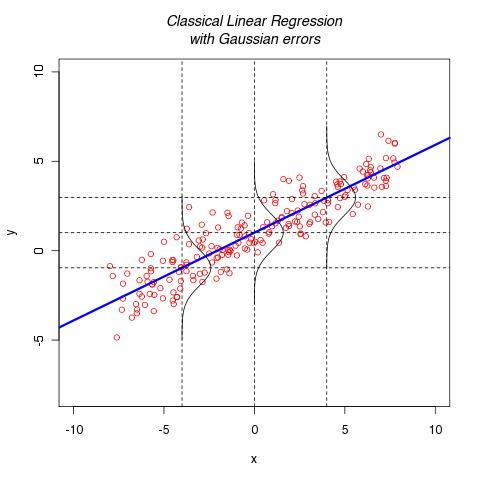 [/caption]
[/caption]
I remember my first encounter to the classical linear regression model with normally distributed errors by Dan Gillen in Stat 120c in spring 2005 at UCI. I mean, I've done linear regression in high school science classes: you get a best fit line out of your data. In the stat class, I remember seeing data drawn on the board and a line going through the data. To illustrate the normally distributed errors, he drew sideway normal curves.
I liked this presentation of the classical model. How do convey this in R? I couldn't find a package for it (searched for an hour I think). I always search for packages because I always think others could do a better job at it than me and because I don't like to re-invent things, even though it requires less time for me to do it myself in many cases. Well, here it is, written by me in R:
```{r}
sideways.dnorm <- function(where.x, where.y, values=seq(-4,4,.1), magnify=4, ...){
###... consists of mean, sd, and log passed to dnorm function
dens <- dnorm(x=values, ...)
x <- where.x + dens * magnify
y <- where.y + values
return(cbind(x,y))
}
temp <- sideways.dnorm(where.x=3, where.y=3, values=seq(-2,2,.1))
plot(temp)
##jpeg("LinearRegressionNormalErrors.jpeg")
pdf("LinearRegressionNormalErros.pdf")
set.seed(123)
n <- 200
x <- runif(n, -8,8)
y <- 1 + .5*x + rnorm(n, sd=1)
fit <- lm(y~x)
plot(x,y, xlim=c(-10,10), ylim=c(-8, 10)
, main="Classical Linear Regression\nwith Gaussian errors"
##, main=expression(paste("Classical Linear Regression\nwith N(0,", sigma\^2, ") errors", sep=''))
, col="red"
)
abline(fit, lwd=3, col="blue")
where.normal.x <- c(-4,0,4)
for(i in 1:length(where.normal.x)){
where.x <- where.normal.x[i]
where.y <- predict(fit, newdata=data.frame(x=where.x))
xy <- sideways.dnorm(where.x=where.x, where.y=where.y, magnify=4)
lines(xy)
abline(h=where.y, lty=2)
abline(v=where.x, lty=2)
}
dev.off()
```